In addition, tissue autofluorescence from endogenous fluorophores (i.e., erythrocytes, collagen, elastin) can result in low signal-to-noise ratios in the imaging data, confounding the detection of weakly expressed antigens. There is also axial and lateral drift during the cyclic imaging process that can lead to misalignment. For example, microscopes are often affected by out-of-focus light and noise from the light source and the camera, resulting in blurry and noisy images.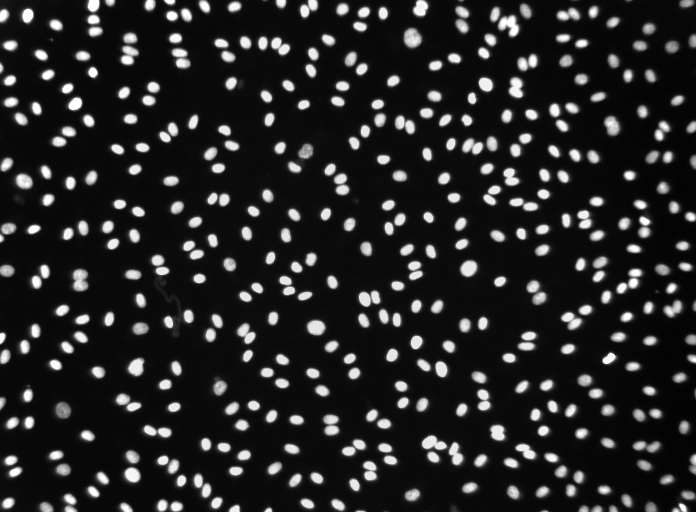 
To view these marker images, it is necessary to generate montages from individual tiles and to align these montages from different cycles to create a high-dimensional imaging dataset also called hyperstack (x, y, channel, cycle).Īt this scale, creation of high-dimensional imaging datasets is challenging. The raw imaging data typically consists of multiple tile scans (i.e., fields of view) over a large tissue region across many cycles with 2 to 4 channel images per tile and multiple z planes per image volume. Moreover, with high-dimensional imaging datasets on the order of terabytes now routinely generated, efficient and accurate image processing is imperative for reliable and reproducible downstream analysis.Īlthough different highly multiplexed fluorescence imaging techniques rely on different modes of antibody tagging (e.g., fluorophores, DNA oligonucleotide barcodes) and different tissue handling protocols, they all involve iterative, multicycle image acquisition using conventional fluorescence microscopes. Given the complexity and time-consuming nature of these experiments, it is necessary to quickly examine the acquired and processed images from the first few cycles as a data sanity check and to allow optimization of experimental conditions (e.g., exposure time) before additional experiments are performed. These multiplexing techniques typically use cyclic immunostaining and imaging to circumvent the spectral limitation of conventional microscopes. Over the past decade, several highly multiplexed fluorescence imaging techniques, including MxIF, t-C圜IF, and CODEX, have been developed to enable the imaging of over 50 antigens in a single tissue section ( 2). High-dimensional, single-cell imaging yields multiscale biological information from the molecular to the cellular to the tissue level to enable the inference of biological mechanisms and to drive therapeutic and diagnostic development ( 1). RAPID reduces data processing time and artifacts and improves image contrast and signal-to-noise compared to our previous image processing pipeline, thus providing a useful tool for accurate and robust analysis of large-scale, multiplexed, fluorescence imaging data. Incorporation of an open source CUDA-driven, GPU-assisted deconvolution produced results similar to fee-based commercial software. RAPID deconvolves large-scale, high-dimensional fluorescence imaging data, stitches and registers images with axial and lateral drift correction, and minimizes tissue autofluorescence such as that introduced by erythrocytes.  
Here, we describe RAPID, a Real-time, GPU-Accelerated Parallelized Image processing software for large-scale multiplexed fluorescence microscopy Data. Fast and accurate processing of these large-scale, high-dimensional imaging data is essential to ensure reliable segmentation and identification of cell types and for characterization of cellular neighborhoods and inference of mechanistic insights. With ever-increasing numbers of antigens, region sizes, and sample sizes, multiplexed fluorescence imaging experiments routinely produce terabytes of data. Highly multiplexed, single-cell imaging has revolutionized our understanding of spatial cellular interactions associated with health and disease.
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
